Research: Dietary differences among individuals with different genes and coat colours gives insight into the maintenance of the Spirit bears among black bear populations
A new study has used a chemical technique (stable isotope analysis) to compare the diets between phenotypes (black and white) and genotypes (0, 1 or 2 copies of the white-causing gene, of which only the 2 copy combination codes for white fur) of coastal black bears.
A new study published by a team of researchers, led by Dr. Christina Service, provides insight into the mechanisms that support the rare white-coated morph of black bear known as the Spirit bear.
The paper, “Intrapopulation foraging niche variation between phenotypes and genotypes of Spirit bear populations,” was published in the open-access journal Ecology and Evolution.
The researchers used stable isotope analysis to compare the diets – or foraging niche – between phenotypes of coastal black bear (white versus black coat), as well as genotypes within the black-coated bears (heterozygotes and 2 versions of the homozygotes). Their results support the hypothesis that the white-coated Spirit bears may have a selective advantage while hunting salmon, as they showed a higher amount of marine-based isotopes found in salmon present during the fall foraging season.
The researchers also found that the heterozygote black-coated bears (which carry one copy of the gene that produces white-coated bears) also show modest divergence in salmon diet compared to the homozygotes, which carry no copies of the white-coding gene. This could help explain the continued selection for the rare white-coated morph.
Abstract
Foraging niche variation within a species can contribute to the maintenance of phenotypic diversity. The multiniche model posits that phenotypes occupying different niches can contribute to the maintenance of balanced polymorphisms. Using coastal populations of black bears (Ursus americanus kermodei) from British Columbia, Canada, we examined potential foraging niche divergence between phenotypes (black and white “Spirit” coat color) and between genotypes (black‐coated homozygote and heterozygous). We applied the Bayesian multivariate models, with biotracers of diet (δ13C and δ15N) together comprising the response variable, to draw inference about foraging niche variation. Variance–covariance matrices from multivariate linear mixed‐effect models were visualized as the Bayesian standard ellipses in δ13C and δ15N isotopic space to assess potential seasonal and annual niche variation between phenotypes and genotypes. We did not detect a difference in annual isotopic foraging niche area in comparisons between genotypes or phenotypes. Consistent with previous field experimental and isotopic analyses, however, we found that white phenotype Spirit bears were modestly more enriched in δ15N during the fall foraging season, though with our modest sample sizes these results were not significant. Although also not statistically significant, variation in isotopic niches between genotypes revealed that heterozygotes were moderately more enriched in δ13C along hair segments grown during fall foraging compared with black‐coated homozygotes. To the extent to which the pattern of elevated δ15N and δ13C may signal the consumption of salmon (Oncorhynchus spp.), as well as the influence of salmon consumption on reproductive fitness, these results suggest that black‐coated heterozygotes could have a minor selective advantage in the fall compared with black‐coated homozygotes. More broadly, our multivariate approach, coupled with knowledge of genetic variation underlying a polymorphic trait, provides new insight into the potential role of a multiniche mechanism in maintaining this rare morph of conservation priority in Canada’s Great Bear Rainforest and could offer new understanding into polymorphisms in other systems.
Citation
Service, C. N., T. Ingram, T. E. Reimchen, C. T. Darimont. 2021. Intrapopulation foraging niche variation between phenotypes and genotypes of Spirit bear populations. Ecology and Evolution. DOI: https://doi.org/10.1002/ece3.7276
Select figures
Figure 1
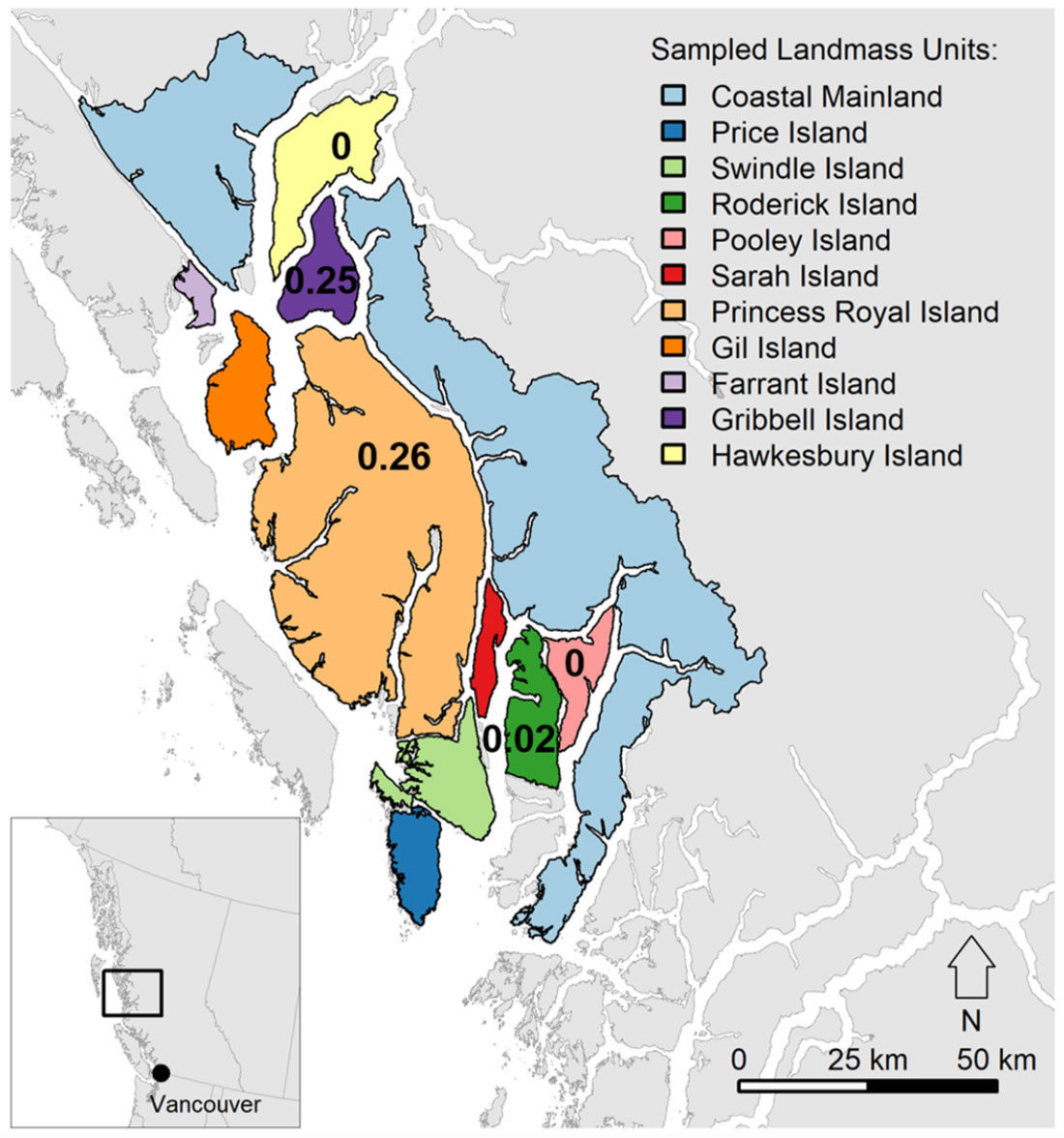
Figure 2

Affiliations
- Department of Geography, University of Victoria, Victoria, BC, Canada
- Raincoast Conservation Foundation, Sidney, BC, Canada
- Kitasoo Xai’xais Stewardship Authority, Kitasoo/Xai’xais First Nation, Klemtu, BC, Canada
- Department of Zoology, University of Otago, Dunedin, New Zealand
- Department of Biology, University of Victoria, Victoria, BC, Canada









